By: Dr Catrin Goebel
Introduction
The detection of the abuse of synthetic insulins by doping laboratories is likely to become a routine requirement. The World Anti-Doping Authority (WADA) code normally requires the use of mass spectrometry to identify prohibited drugs but peptide hormones are currently excluded because of the difficulty of obtaining mass spectra from such large molecules at very low physiological concentrations. Recent developments in applying mass spectrometry to proteomics means that it is becoming feasible for doping laboratories to routinely apply such methodology to detect and confirm the abuse of peptide hormones. The methodology to detect and confirm the abuse of peptide hormones by mass spectrometry is preferred to the current use of immunoassays or other immuno-reactive techniques. Insulin is a clear example of how both endogenous insulin and its synthetic analogues can give a positive result with some immunological assays but mass spectral analysis can easily distinguish between them. The use of all types of insulin is prohibited by non-diabetic athletes but it is desirable if possible to identify which form of insulin has been used.
To use techniques such as liquid chromatography-mass spectrometry (LC-MS) to detect synthetic insulins requires that they be extracted and concentrated prior to LCMS analysis. This requires at least one evaporation of a solution with high water content. It has been found for insulin-containing solutions that a vacuum concentrator is superior to nitrogen evaporation as it is much faster and gives better recoveries.
Insulin is a peptide hormone consisting of two peptide chains that are cross-linked by two disulfide bridges (Figure 1). The two peptide chains are denoted as chain-A and chain-B. Five synthetic insulins were studied, Apidra (Aventis), Humalog (Eli Lilly &
Company), Lantus (Aventis), Levemir (Novo Nordisk Pharmaceuticals Pty. Ltd.) and Novorapid (Novo Nordisk Pharmaceuticals Pty. Ltd.), for the analysis in serum samples by liquid chromatography tandem mass spectrometry (LC/MS/MS) with electrospray ionisation. The synthetic insulins can be differentiated from endogenous human insulin because their amino acid sequence has been modified. The simplest modification is the swap of positions for the proline and lysine from human insulin to Humalog (B-chain residue 28 and 29), as highlighted in Figure 1. Humalog is then the hardest to differentiate by LC/MS/MS as the observed precursor molecular ions ([M + nH]+n) are the same but for the product ions there is a clear distinction. The m/z 217 is observed for Humalog fragment y2 (ProThr, B29B30) while for human insulin the m/z 226 is observed representing y3 – y1 (ProLys, B28B29).
Figure 1: Insulin amino acid sequence with disulfide bonds. The yellow highlighted amino acids indicate the a change from human insulin

Apidra, a rapid acting human insulin analog, has asparagine at position B3 replaced by lysine and lysine at position B29 has been replaced by glutamic acid (Figure 1). Three modifications have been made for the insulin analog Lantus: glycine replace asparagine at A21 and two arginine amino acids are added to the COOH-terminal of the B chain. Threonine has been omitted from Levemir and a C14 fatty acid chain has been attached to the B29 amino acid, resulting in a long-acting analog. Novorapid has the amino acid proline at B28 replaced with aspartic acid (Figure 1), which introduces an additional negative charge within the insulin molecule causing the rapid action of the product.
Apidra, Lantus, Levemir and Novorapid are easily distinguished by LC/MS/MS analysis as each has distinct precursor ions because of the variation in molecular weight as well as distinct product ions.
Extraction, Concentration and Analysis
Insulin was extracted from 2 mL serum samples using immunoaffinity chromatography columns followed by a solid phase extraction. The final volume of the purified samples is 1.2 mL in aqueous 2% acetic acid with 50% acetonitrile. The extraction technique implemented within ASDTL had been developed by 1Thevis et al.
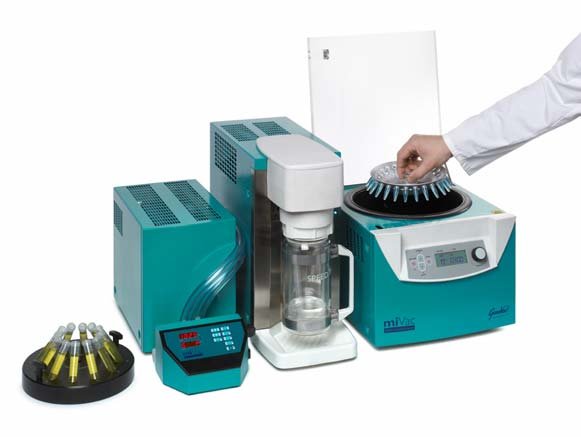
The extracts were dried using either a Turbovap (Zymark) with nitrogen gas supply and water bath set to 35ºC or a miVac DUO with Quattro pump (Genevac – Figure 2). The samples were analysed on a 4000 Q TRAP (Applied Biosystems) with a gradient separation (Agilent 1100, CapPump Binary) using a XBridge Shield RP18 3.5μm 1.0 x 150mm column. Solvent A consisted of 95% H2O 5% acetonitrile and solvent B consisted of 90% acetonitrile 10% H2O both A and B had 0.2% formic acid. The gradient was constant from 0 to 2 minutes at 85% A, then reduced to 32.5% A to 14 minutes, solvent A increased back to 85% at 15 minutes and equilibrated the column at 85% A till 20 minutes.
Figure 2: miVac Duo concentrator system

Results
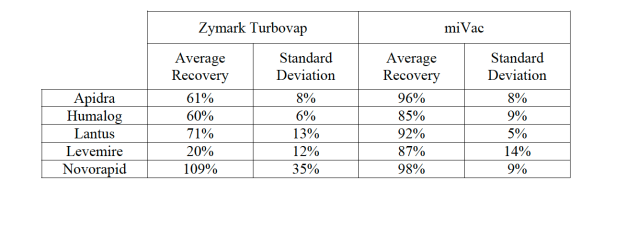
The samples would require up to 6 hours drying time using the Turbovap which resulted in the preparation taking two days before samples would be ready for analysis by LC/MS/MS. In comparison the miVac DUO dried samples in 2 hours using the alcohol method with heating set to 40ºC. The samples could then be extracted and ready for analysis within a single day. Furthermore the recoveries and recovery reproducibility of all five insulin analogs, Apidra, Humalog, Lantus,
Levemire and Novorapid, were significantly improved as shown in Figure 3.
Figure 3: Recovery and standard deviation for six replicate extractions by immunoaffinity chromatography
Summary
The use of the miVac DUO with Quattro DUO pump significantly improved the method for the detection of synthetic insulins in serum by being both much faster and by giving better and more consistent recoveries than obtained with nitrogen evaporation.
Reference
Thevis M, Thomas A, Delahaut P, Bosseloir A, Schänzer W. Qualitative determination of synthetic analogues of insulin in human plasma by immunoaffinity purification and liquid chromatography-tandem mass spectrometry for doping control purposes. Anal Chem. 2005 Jun 1;77(11):3579-85.
